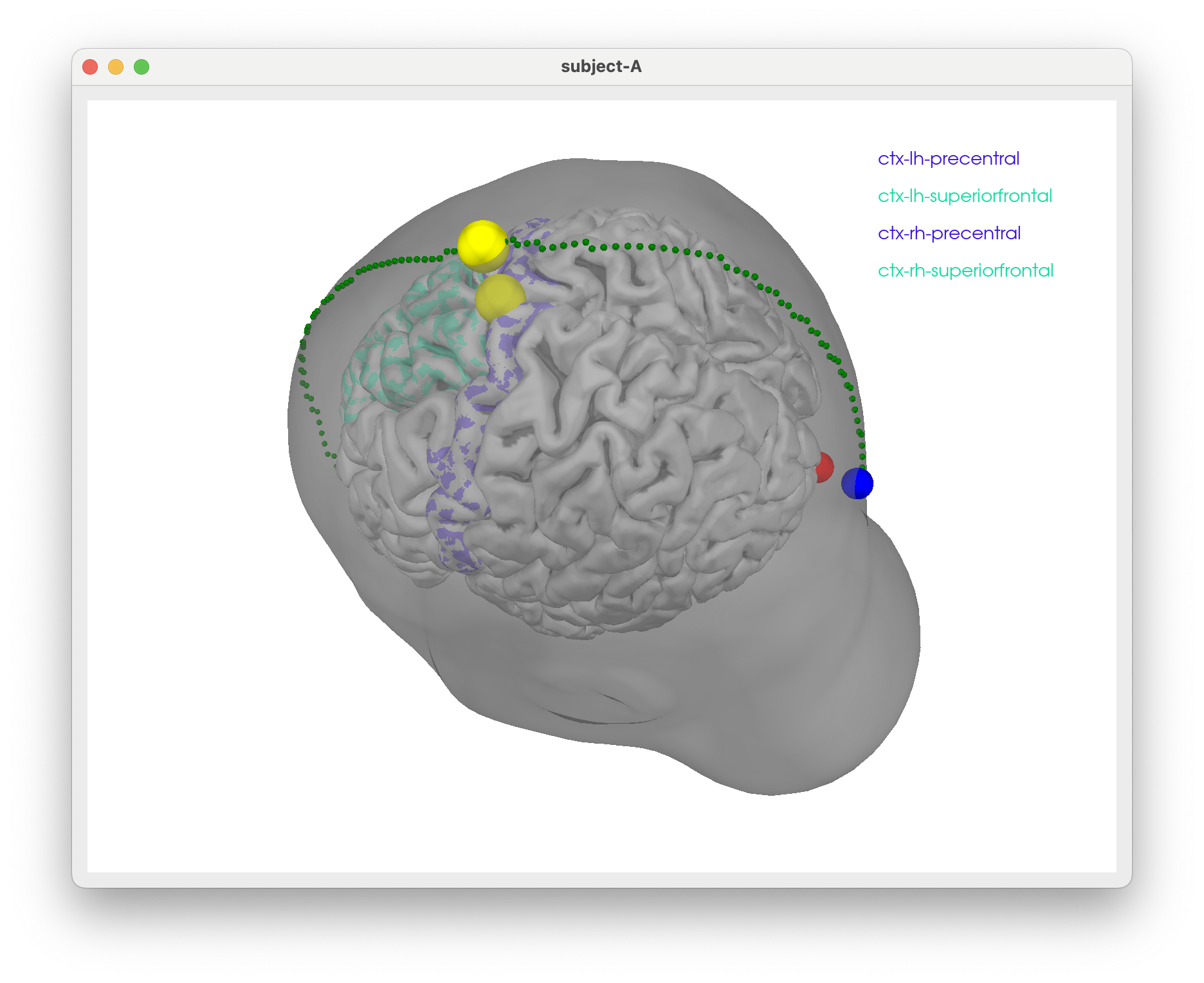
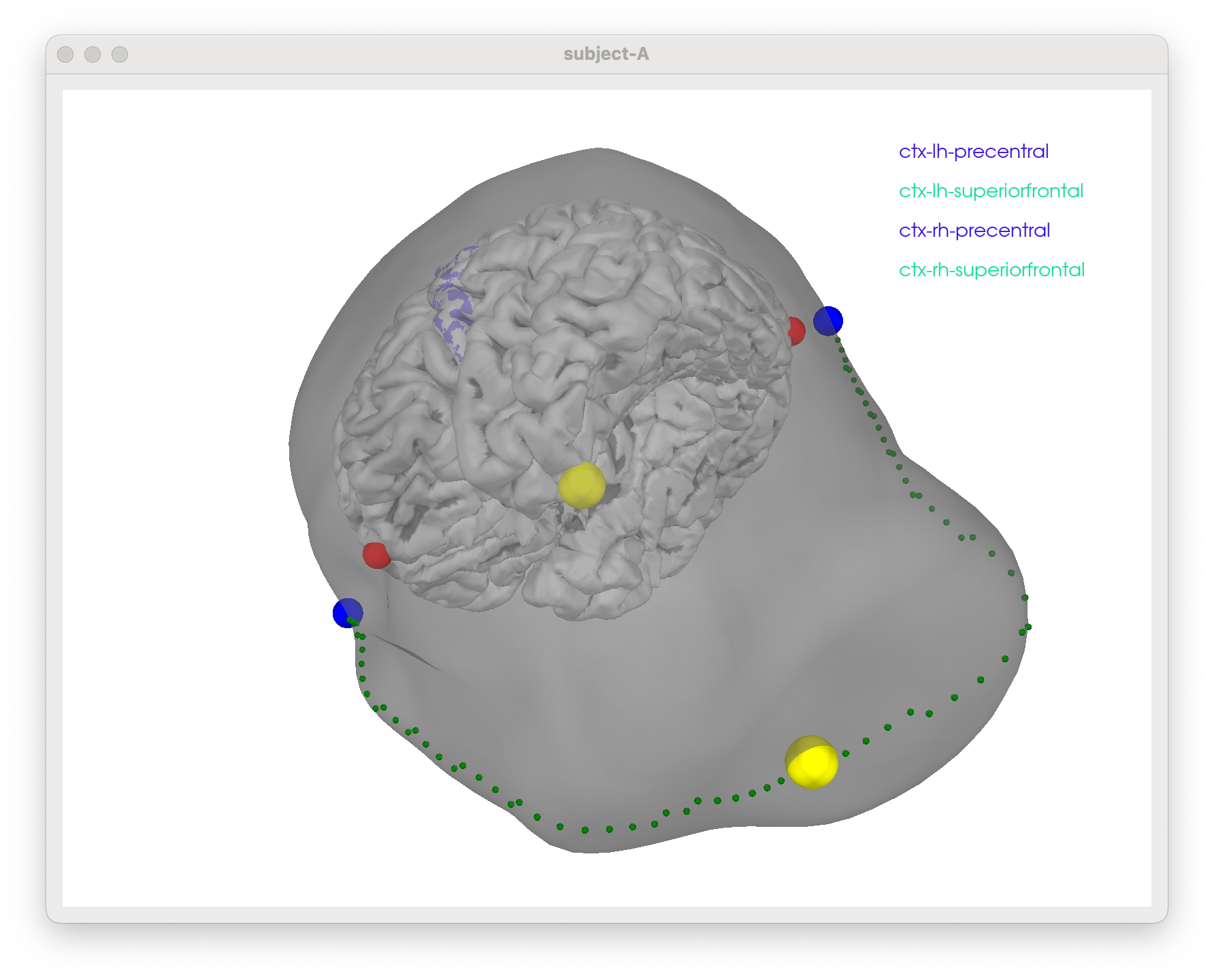
tms-head-recon
Some freesurfer magic and MNE viz
Some notes to figure out how to show TMS stim sites scalp/skull surfaces w/ promixity to particular regions in the brain
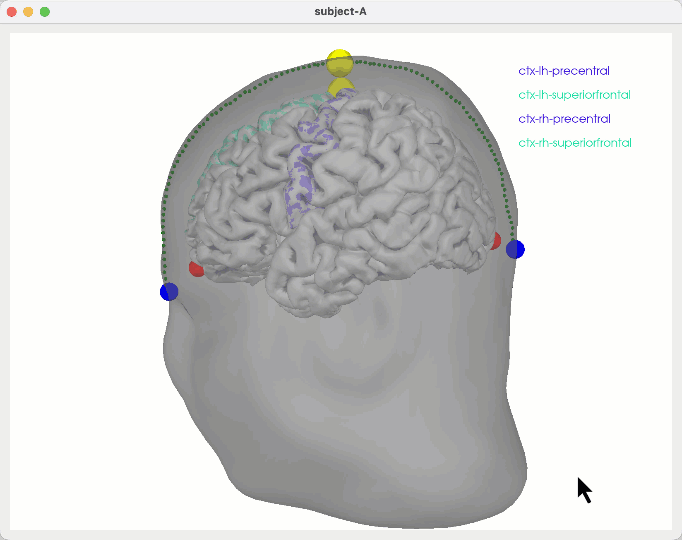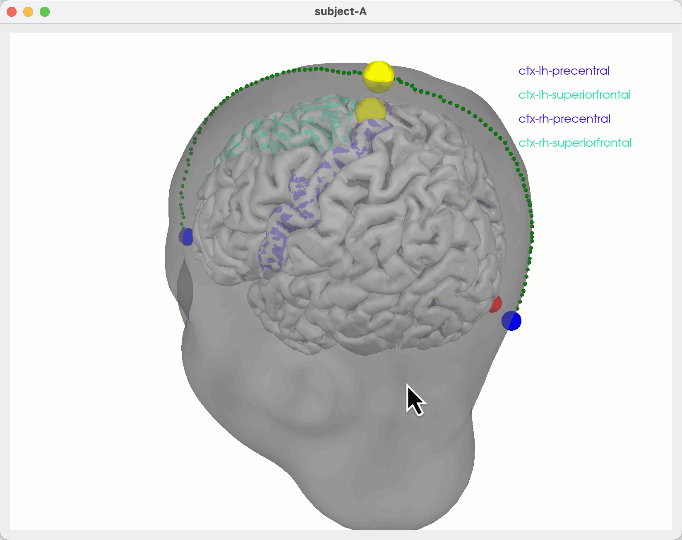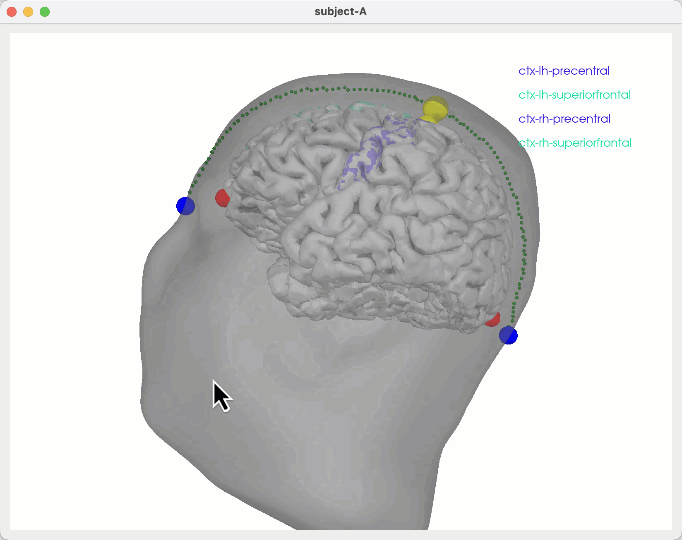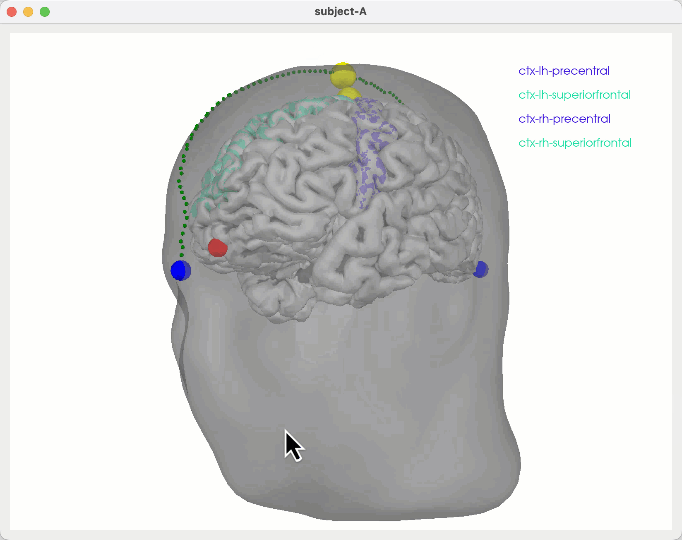
General ideas
- use
freesurferto generate scalp/skull surfaces from T1 MRI - use
mnetools to generate BEM surfaces from freesurfer outputs - use
mneto visualize TMS coil positions relative to scalp/skull/brain surfaces and/ormatlabwith specific freesurfer visualization tools
-
Assuming you have run freesurfer’s
recon-allon your T1 MRI to generate the necessary surfaces and have installedmnetools on your machine + havematlabknocking around 😄! -
Use
mne watershed_bemto generate scalp and skull surfaces
SUBNAME=subject-A
mne watershed_bem -s ${SUBNAME}
and visualise:
SUBNAME=subject-A
mne freeview_bem_surfaces -s ${SUBNAME}
This will allow you to get some nice screengrabs of skull / skin surfaces. Etc.
I also used this to approximate the inion/nasion points on the scalp surface for Cz locating. See code to check if that makes sense
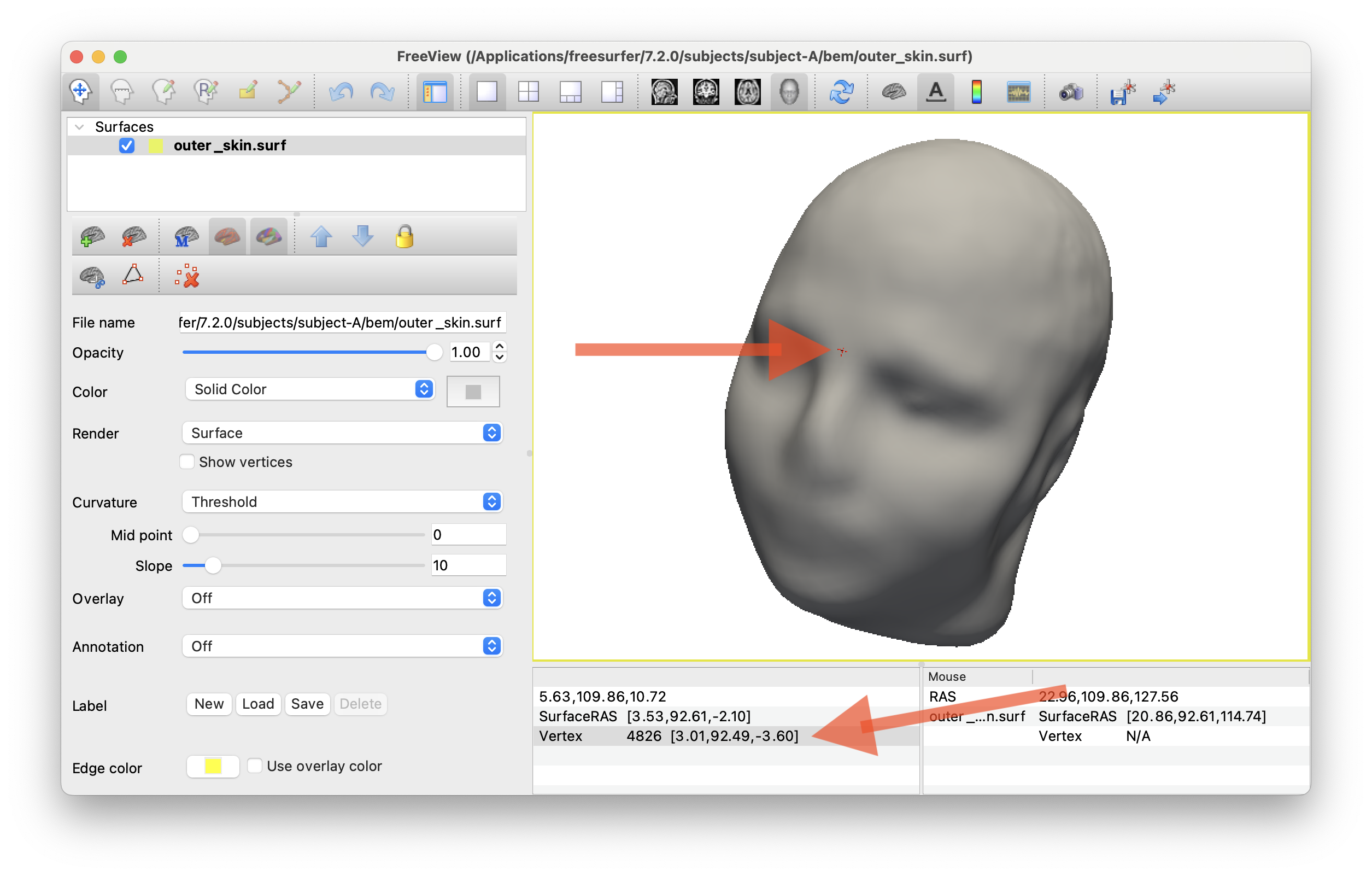
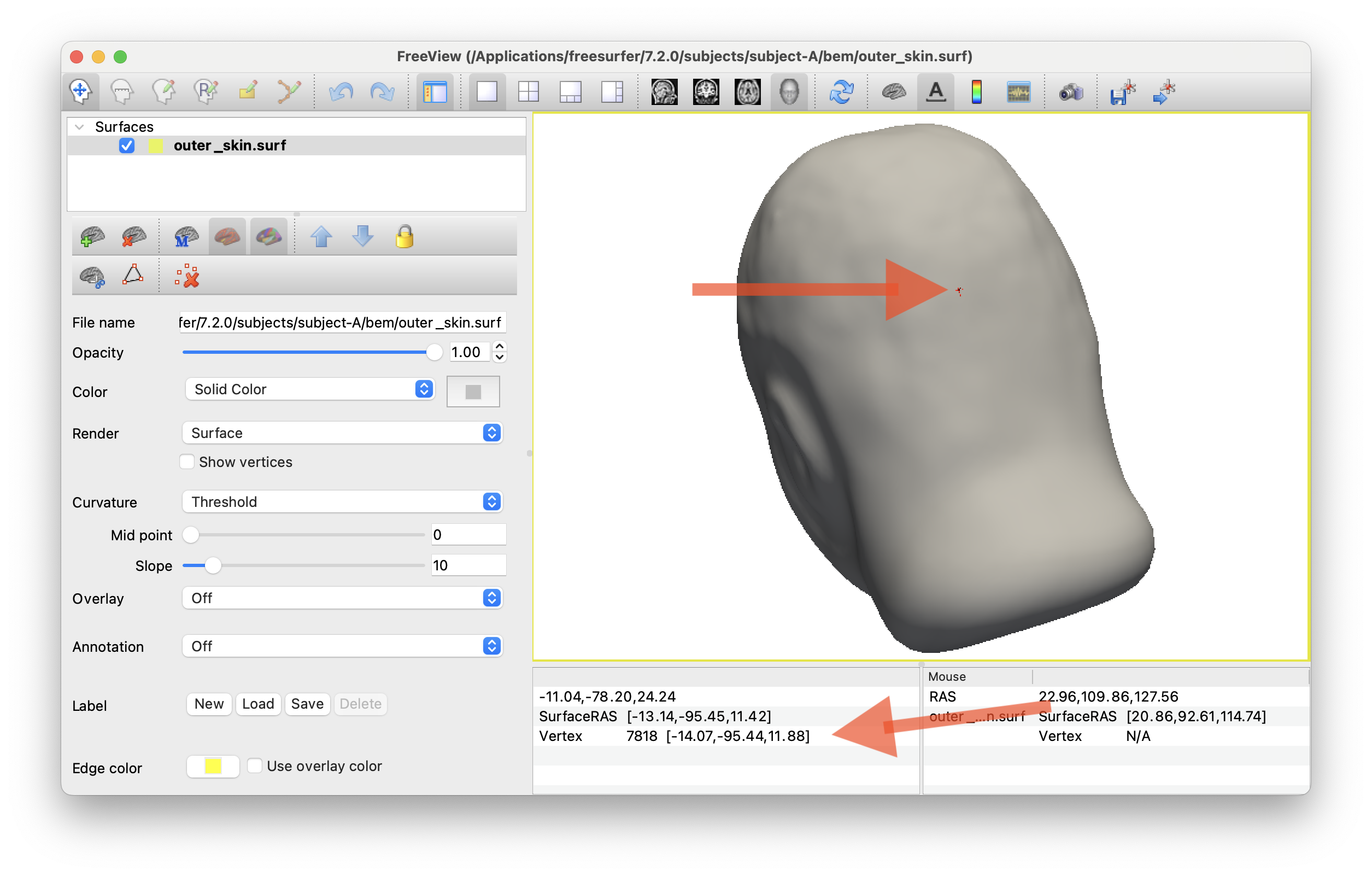
Visualizing TMS coil positions w/ CZ etc.
Probably easiest to do this in python using mne.viz. Here’s some example code to get you started h/t prompts from Claude in Visual Studio Code and the documentation on the MNE site.
- Make sure the right python environment is activated with
mneinstalled
# if you used the MNE installer, something like:
conda activate /Applications/MNE-Python/1.11.0_0/.mne-python
You should be able to run the interative viz code below and see the scalp/skull surfaces along with fiducials and Cz of montage, if I got that correct. Inspect code at to see details.
# make sure right python!
conda activate /Applications/MNE-Python/1.11.0_0/.mne-python
# then run
python ./create_montage_viz.py

Etc. etc…
Penalty for particular paths
- code includes a penalty for paths that deviate from sagittal plane
x_devis penalized. - to avoid the shorted path hugging the surface inferiorly (when skull/scalp surfaces are relatively clipped inferiorly), a penalty for low z values (when
z_posis negative) is included.
# default behaviour PREFER upwards paths
python create_montage_viz.py # default behaviour
python create_montage_viz.py up # default behaviour
# as a reality check you can also try to penalise upwards paths by PREFERing downwards paths
python create_montage_viz.py down